


 النبات
النبات
 الحيوان
الحيوان
 الأحياء المجهرية
الأحياء المجهرية
 علم الأمراض
علم الأمراض
 التقانة الإحيائية
التقانة الإحيائية
 التقنية الحيوية المكروبية
التقنية الحيوية المكروبية
 التقنية الحياتية النانوية
التقنية الحياتية النانوية
 علم الأجنة
علم الأجنة
 الأحياء الجزيئي
الأحياء الجزيئي
 علم وظائف الأعضاء
علم وظائف الأعضاء
 الغدد
الغدد
 المضادات الحيوية
المضادات الحيوية|
Read More
Date: 24-12-2015
Date: 29-11-2015
Date: 15-5-2016
|
His Operon
Energy equivalent to about 41 ATP molecules is required to synthesize one molecule of the amino acid histidine (1). The considerable metabolic cost of histidine biosynthesis presumably accounts for the evolution of multiple strategies to regulate the rate of synthesis of the amino acid in response to environmental changes. Checkpoints regulate both the flow of intermediates through the biosynthetic pathway and the amounts of histidine-biosynthetic enzymes present. Expression of the genes for
these enzymes is regulated in bacterial cells by mechanisms that are both general (metabolic regulation, elongation control) and specific (attenuation control, segmental stabilization of the distal part of the messenger RNA).
1.Structural organization of the operon
In Escherichia coli and Salmonella typhimurium the enzymes responsible for the biosynthesis of histidine are encoded by eight genes tightly clustered in a single, large operon (his operon). In both species, transcription produces a single polycistronic mRNA about 7300 nucleotides long, extending from a primary promoter (hisp1) to a Rho-independent terminator. Two weak internal promoters, hisp2 and hisp3, are located within the hisC and hisF genes, respectively. The structural organization of the operon is essentially the same in the two species, and in both the translational stop codon of each cistron overlaps the translational initiation codon of the downstream cistron (2). This organization allows ribosomes to initiate the translation of a new cistron without moving away from the mRNA after terminating translation of the preceding one. Such a translational coupling mechanism probably guarantees equimolar synthesis of the corresponding gene products (3).
2. Control of transcription initiation and elongation
Transcription of the his operon is about four-fold more efficient in bacteria growing in minimal-glucose medium than when growing in rich medium. This form of control, called metabolic regulation, adjusts the expression of the operon to the amino acid supply in the cell. It is mediated by the “alarmone” guanosine 5′-diphosphate 3′-diphosphate (ppGpp), which is the effector of the stringent response. The alarmone regulates the his operon positively by stimulating the primary promoter hisp1 under conditions of moderate amino acid starvation (4. (
In addition to this general metabolic control, his operon transcription is specifically regulated by attenuation of transcription, a mechanism in which a regulatory element, located upstream of the first structural gene of the cluster, modulates the level of expression of the histidine biosynthetic enzymes in response to the intracellular levels of charged histidyl-transfer RNA, His-tRNAHis (5). The his-specific regulatory element is transcribed in a 180-nucleotide RNA leader, which exhibits two prominent features: (i) a 16-residue coding sequence including seven consecutive codons specifying histidine, and (ii) overlapping regions of dyad symmetry capable of folding into mutually exclusive, alternative secondary structures that signal either transcription termination or antitermination (6, 7). Six RNA segments are involved in base pairing (Fig. 1 A to F) and the stem-loop structure formed by the E and F RNA regions, plus the adjacent run of uridylate residues, constitutes the attenuator, a strong Rho-independent transcription terminator (Fig 1). Translational control of his operon transcription is determined by ribosome occupancy of the leader RNA, which in turn depends, given the peculiar composition of the his leader peptide, on the availability of His-tRNAHis. High levels of His-tRNAHis allow rapid movement of ribosomes up to the B segment; in this case, formation of the C:D and E:F stem-loop structures will result in premature transcription termination (Fig. 1, Attenuation). In the presence of low levels of charged tRNAHis, ribosomes stall at the consecutive histidine codons of the leader peptide and prevent the A:B pairing by masking the A segment. Base pairing between the B and C and between the D and E RNA regions prevents formation of the attenuator and determines the antitermination conformation (Fig. 1, Transcription). In the case of severe limitation of the intracellular pool of all charged tRNAs, translation of the leader peptide fails to initiate: under these conditions, the A:B, C:D and E:F stem-loop structures form sequentially, producing a strong transcription termination (Fig. 1, Superattenuation). RNA polymerase pauses after synthesis of the first RNA hairpin (A:B). This pausing is believed to synchronize transcription and translation of the leader region by halting the elongating RNA polymerase until a ribosome starts translation of the leader peptide (8). The pause hairpin (Fig. 1) is the only portion of the structure thought to form when RNA polymerase resides at the pause site.
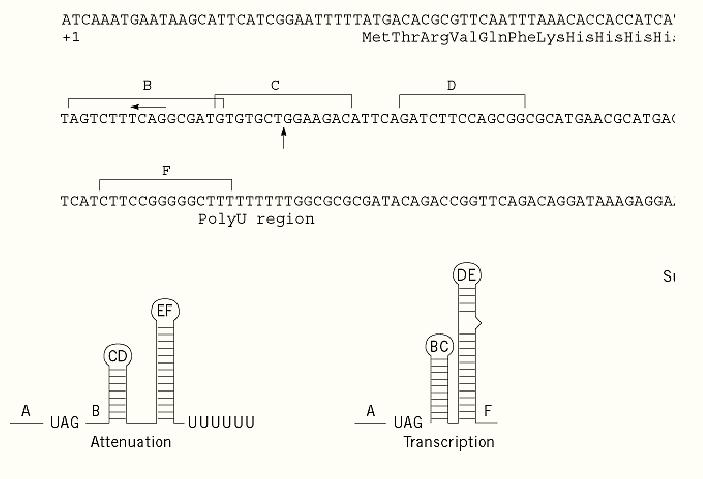
Figure 1. Regulation of translation of the his operon messenger RNA.
Because the absolute amount of charged tRNAHis controls the level of his attenuation (5), mutants exhibiting high his operon expression contain defects in tRNAHis biosynthesis, aminoacylation with histidine, or tRNA His modification and processing. The hisR gene encodes the single cellular tRNAHis; and mutations in the hisR promoter reduce the total cellular content of tRNAHis molecules by about 50% and thereby cause increased readthrough transcription of the his attenuator (9). The hisS gene encodes histidyl-aminoacyl tRNA synthetase, which aminoacylates tRNAHis molecules with histidine. Mutations that lower the activity of the histidyl-tRNA synthetase or decrease the enzyme's affinity for histidine, tRNAHis, or ATP, affect the level of his attenuation by reducing the percentage of tRNAHis molecules charged with histidine (10). The hisT gene encodes pseudouridine synthase I, which catalyzes the formation of pseudouridine residues in the anticodon region of several tRNA species, including tRNAHis. Although the undermodified tRNAHis molecules are charged with histidine to the same extent as in wild-type strains, transcription termination at the his attenuator is greatly decreased, because the slow rate of translation of the consecutive histidine codons causes stalling of ribosomes (11. (
The overall contribution of the internal promoter hisp2 to the expression of the distal genes of the operon is negligible when transcription proceeds from hisp1, because hisp2 is inhibited by transcription readthrough, a phenomenon known as promoter occlusion (12). hisp2 is also subjected to metabolic regulation, although to a lesser extent than hisp1.
Elongation of the his-mRNA is modulated by a non-specific mechanism operating at the level of intracistronic transcription termination elements (TTEs) (13). These elements consist of cytosine-rich and guanosine-poor RNA regions and are the binding-activation sites of the transcription-termination Rho factor, which is responsible for polar effects in polycistronic operons (14). A premature arrest of translation, produced by nonsense mutations, favors the binding of Rho to the TTE on the nascent transcript. The subsequent interaction of Rho with the elongating RNA polymerase causes a premature release of transcripts. Polarity results, with reduced expression of the genes located downstream from the TTE.
3. Decay and Segmental Stabilization of the his-mRNA
The primary 7300-nucleotide his-mRNA has a half-life of about 3 minutes in cells growing in minimal-glucose medium and is degraded in a net 5′ → 3′ direction. Three major processed species, 6300, 5000, and 3900 nucleotides long, encompassing the last seven, six, and five cistrons, respectively, are generated in the decay process (12). The 6300- and the 5000-nucleotide RNAs, which have half-lives of 5 and 6 minutes, respectively, have heterogeneous 5′ ends generated by ribonuclease E cleavage. The 3900-nucleotide processed RNA species has a unique 5′ end and an uncommon stability, having a half-life of about 15 minutes. This RNA species is generated by specific processing events requiring sequential cleavages by two different endonucleases. RNase E triggers the process by cleaving at a major target site located in the hisC cistron, 620 nucleotides upstream of the 5′ end of the processed species. Subsequently, ribonuclease P cleaves the processed RNA species generated by RNase E at a site located 76 nucleotides upstream of the start codon of the hisB cistron, producing the mature 5′ end. The observation that the RNase P-catalyzed reaction requires the presence of ribosomes suggests that translation of the hisB cistron might favor formation of the structure recognized by RNase P (15).
References
1. M. Brenner and B. N. Ames (1971) In Metabolic Pathways (H. J. Vogel, ed.), Academic Press, Inc., N.Y., vol. 5, pp. 349–387.
2. M. S. Carlomagno, L. Chiariotti, P. Alifano, A. G. Nappo, and C. B. Bruni (1988) J. Mol. Biol. 203, 585-606.
3. C. Yanofsky, T. Platt, I. P. Crawford, B. P. Nichols, G. E. Christie, H. Horowitz, M. Van Cleemput, and A. M. Wu (1981) Nucleic Acids Res. 9, 6647–6668.
4. J. C. Stephens, S. W. Artz, and B. N. Ames (1975) Proc. Natl. Acad. Sci. USA 72, 4389–4393.
5.F. Blasi and C. B. Bruni (1981) Curr. Top. Cell. Regul. 19, 1–45.
6.W. M. Barnes (1978) Proc. Natl. Acad. Sci. USA 75, 4281–4285.
7. P. P. Di Nocera, F. Blasi, R. Di Lauro, R. Frunzio, and C. B. Bruni (1978) Proc. Natl. Acad. Sci. USA 75, 4276–4280.
8. C. L. Chan and R. Landick (1989) J. Biol. Chem. 264, 20796–20804.
9. H. M. Johnston, W. M. Barnes, F. G. Chumley, L. Bossi, and J. R. Roth (1980) Proc. Natl. Acad. Sci. USA 77, 508–512.
10. J. A. Lewis and B. N. Ames (1972) J. Mol. Biol. 66, 131–142.
11. C. F. Singer, G. R. Smith, R. Cortese, and B. N. Ames (1972) Nature New Biol. 238, 72–74.
12. P. Alifano, C. Piscitelli, V. Blasi, F. Rivellini, A. G. Nappo, C. B. Bruni, and M. S. Carlomagno (1992) Mol. Microbiol. 6, 787–798.
13. P. Alifano, F. Rivellini, D. Limauro, C. B. Bruni, and M. S. Carlomagno (1991) Cell 64, 553–563.
14. P. Alifano, M. S. Ciampi, A. G. Nappo, C. B. Bruni, and M. S. Carlomagno (1988) Cell 55, 351-360.
15. P. Alifano, F. Rivellini, C. Piscitelli, C. M. Arraiano, C. B. Bruni, and M. S. Carlomagno (1994(Genes Dev. 8, 3021–3031.

|
|
|
|
اكتشاف تأثير صحي مزدوج لتلوث الهواء على البالغين في منتصف العمر
|
|
|
|
|
|
|
زهور برية شائعة لتر ميم الأعصاب التالفة
|
|
|
|
|
|
موكب أهالي كربلاء يستذكر شهادة الإمام الصادق (عليه السلام)
|
|
|
|
العتبة العباسية تستذكر شهادة الإمام الصادق (عليه السلام) بإقامة مجلس عزاء
|
|
|
|
أهالي كربلاء يحيون ذكرى شهادة الإمام الصادق (عليه السلام) في مدينة الكاظمية
|
|
|
|
شعبة مدارس الكفيل النسوية تعقد اجتماعًا تحضيريًّا لوضع الأسئلة الامتحانية
|